
Emre Aksay, Ph.D.
- Associate Professor of Computational Neuroscience in the Institute for Computational Biomedicine
- Director of the Graduate Program in Physiology, Biophysics and Systems Biology
Whitney Pavillion
525 East 68th Street, Room W-820 B
New York, NY 10065
Techniques
- Animal Models (e.g., zebrafish)
- Calcium Imaging
- Electrophysiology (Patch-clamp; whole cell; multi-electrode arrays)
- Fluorescence Microscopy and FRAP
- High Performance Computing
- Machine Learning, AI, Deep Learning
- Mathematical Modeling and Simulations
- Optogenetics
Research Areas
Research Summary:
Dynamics in Neural Systems
We are interested in understanding the molecular, cellular, and circuit mechanisms that give rise to the rich neural dynamics observed in the brain. Neural dynamics, the ongoing time‐varying activity patterns in populations of neurons, are critical for a wide range of motor and cognitive behaviors. To understand neural dynamics, we work at the interface between physics and biology, applying in awake animals an approach that combines molecular‐genetic manipulations, electrophysiology, multi‐photon imaging and perturbation, connectomics, statistical and machine learning, computational modeling, and control theory. Our efforts not only yield basic science insights into neuronal computations and how neurons interact to generate global function, but also help outline therapeutic strategies for dealing with disorders of neural dynamics.
Persistent Neural Activity
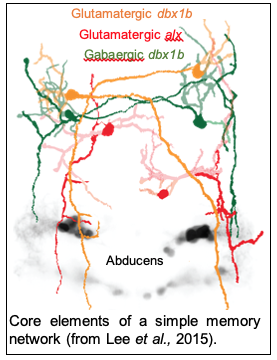 The ability to keep important information in a short-term memory ‘buffer’ is critical for many cognitive, navigational, and motor behaviors. In many settings the biological correlate to short-term memory is persistent neural firing, a sustained change in the level of activity across a network of neurons. In our research on a simple motor memory we have shown that excitatory interactions between neurons in the memory population provide positive feedback that supports persistent firing, inhibitory interactions coordinate the activation of competing subpopulations, and patterns of persistent firing vary with behavioral context. Going forward, we are uncovering the fine-structure of synaptic interactions in the network, how synaptic and dendritic response properties mediate interactions, and the role of different memory patterns in controlling behavior.
The ability to keep important information in a short-term memory ‘buffer’ is critical for many cognitive, navigational, and motor behaviors. In many settings the biological correlate to short-term memory is persistent neural firing, a sustained change in the level of activity across a network of neurons. In our research on a simple motor memory we have shown that excitatory interactions between neurons in the memory population provide positive feedback that supports persistent firing, inhibitory interactions coordinate the activation of competing subpopulations, and patterns of persistent firing vary with behavioral context. Going forward, we are uncovering the fine-structure of synaptic interactions in the network, how synaptic and dendritic response properties mediate interactions, and the role of different memory patterns in controlling behavior.
Plasticity of Neural Dynamics
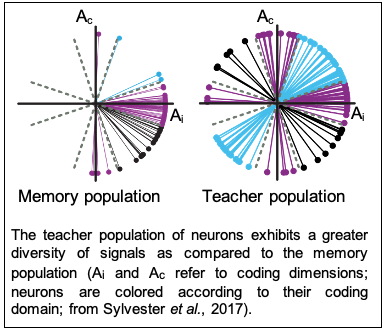
Accurate behavior requires neural dynamics to adapt with changes in the environment or body mechanics. For the motor memory noted above, adaptation in persistent neural firing depends upon interactions between a ‘teacher’ population and the memory population. In our past research we’ve found that in naïve conditions signals in the teacher population are more diverse than those in the memory population by virtue of a form of ‘expansion coding’, leading us to hypothesize that this diversity enables learning of novel patterns of activity in the memory group. We are now investigating what teaching signals are most engaged during learning, the interaction patterns between teacher and memory populations, and the cellular learning rules that generate changes in overall network dynamics.
Ramp-to-Threshold Dynamics
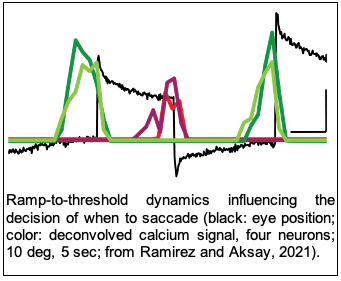
The nervous system must continuously assess evidence and readiness in order to make decisions. Studies have found that activity in some neural populations increases in a ramp-like manner towards a threshold value that is reached at the time a decision is made. We’ve recently found such ramp-to-threshold dynamics in a neural population is associated with deciding when to make a simple spontaneous behavior and demonstrated that perturbing this population disrupts decision-making, thus establishing a causal role for this brain dynamic. Future work is focused on understanding the cellular and circuit mechanisms giving rise to ramp dynamics, the nature of any evidence signals being evaluated, and the role of stochasticity in generating threshold-crossings that trigger decisions.
Epileptiform Dynamics
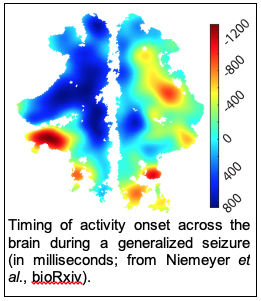 Epilepsy is a disease of aberrant brain dynamics that continues to cause significant disruption to the health and financial security of millions of patients. It is generally agreed that seizures arise due to poor coordination of the activity in excitatory and inhibitory neuronal populations, but the causes of this dysfunction are not well understood. We’ve recently found that clusters of excitatory neurons at distinct initiation sites are disproportionately active during the early phase of seizures. Going forward, we are interested in determining the mechanisms that lead these clusters to form and manipulating the activity of these clusters to control seizure propagation.
Epilepsy is a disease of aberrant brain dynamics that continues to cause significant disruption to the health and financial security of millions of patients. It is generally agreed that seizures arise due to poor coordination of the activity in excitatory and inhibitory neuronal populations, but the causes of this dysfunction are not well understood. We’ve recently found that clusters of excitatory neurons at distinct initiation sites are disproportionately active during the early phase of seizures. Going forward, we are interested in determining the mechanisms that lead these clusters to form and manipulating the activity of these clusters to control seizure propagation.

